Figure 5
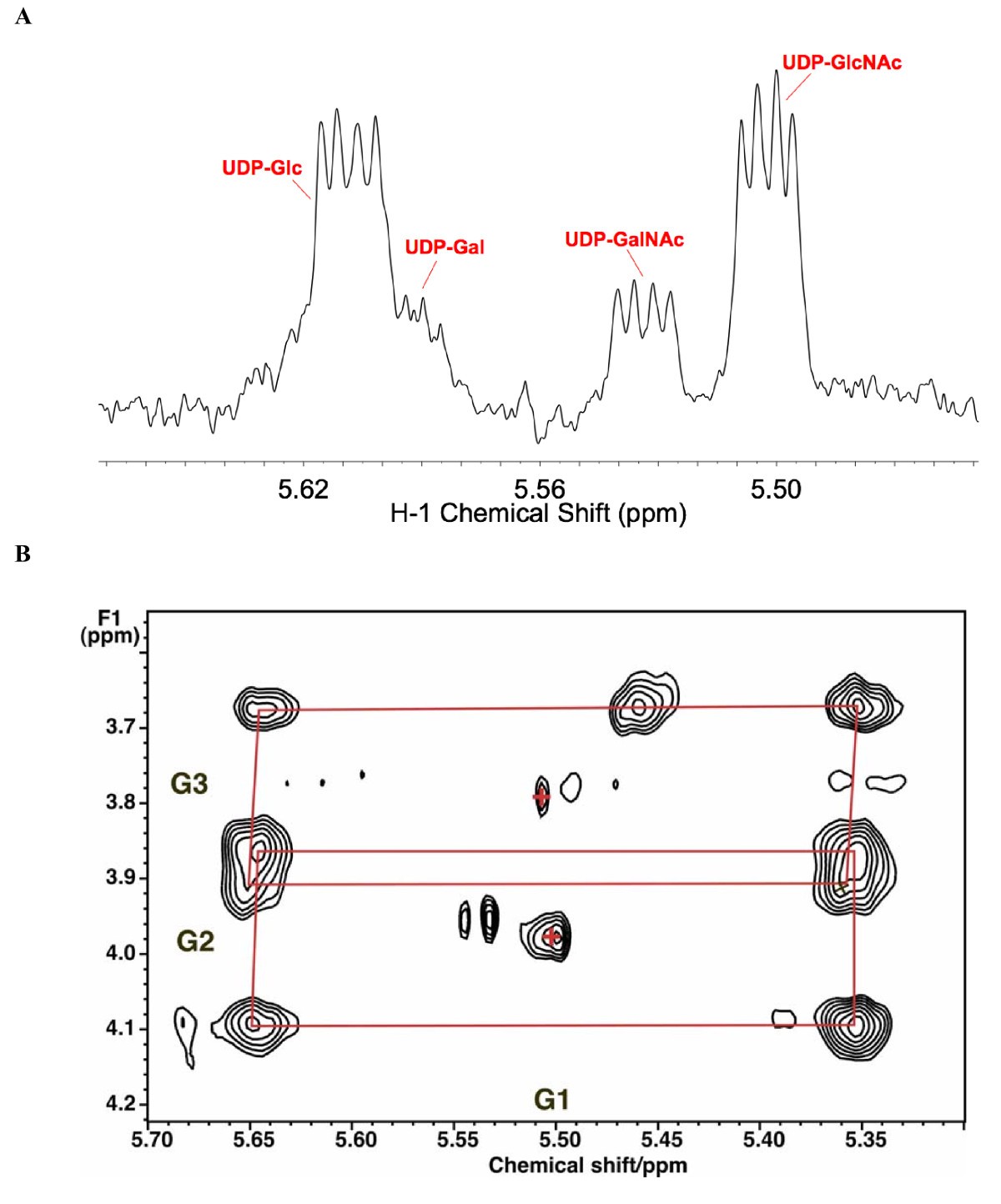
NMR identification of UDP-hexoses. LN3 cells were grown in unlabeled glucose and extracted as described in the Methods section. One-dimensional 1H NMR spectra were recorded at 20°C, 800 MHz. Two-dimensional 1H total correlation spectroscopy (TOCSY) spectra were recorded at 600 MHz using a mixing time of 50 ms with a spin lock field strength of 8 kHz. (a) One-dimensional NMR spectrum. The sugar anomeric region shows several resonances that are double doublets, that is, a single proton scalar coupled to two different spin 1/2 nuclei. By comparison with standard spectra, these have been assigned to the glucose H1 of UDP-glucose, UDP-galactose (UDP-Gal), UDP-GalNAc and UDP-GlcNAc by 1H and 13C chemical shifts, splitting patterns, 13C labeling and two-dimensional TOCSY crosspeak patterns as described in the text. (b) Anomeric region of example TOCSY spectrum showing 13C satellites of glucose H1-H3 of UDP-GlcNAc due to incorporation of [U-13C]-glucose. The satellite crosspeaks (denoted by red rectangles) represent the covalent linkages of H1 to H2 and H3 of the glucose moiety (labeled respectively as G1, G2 and G3), which were essentially completely labeled at 48 h. Crosses denote the crosspeaks of protons attached to 12C.
